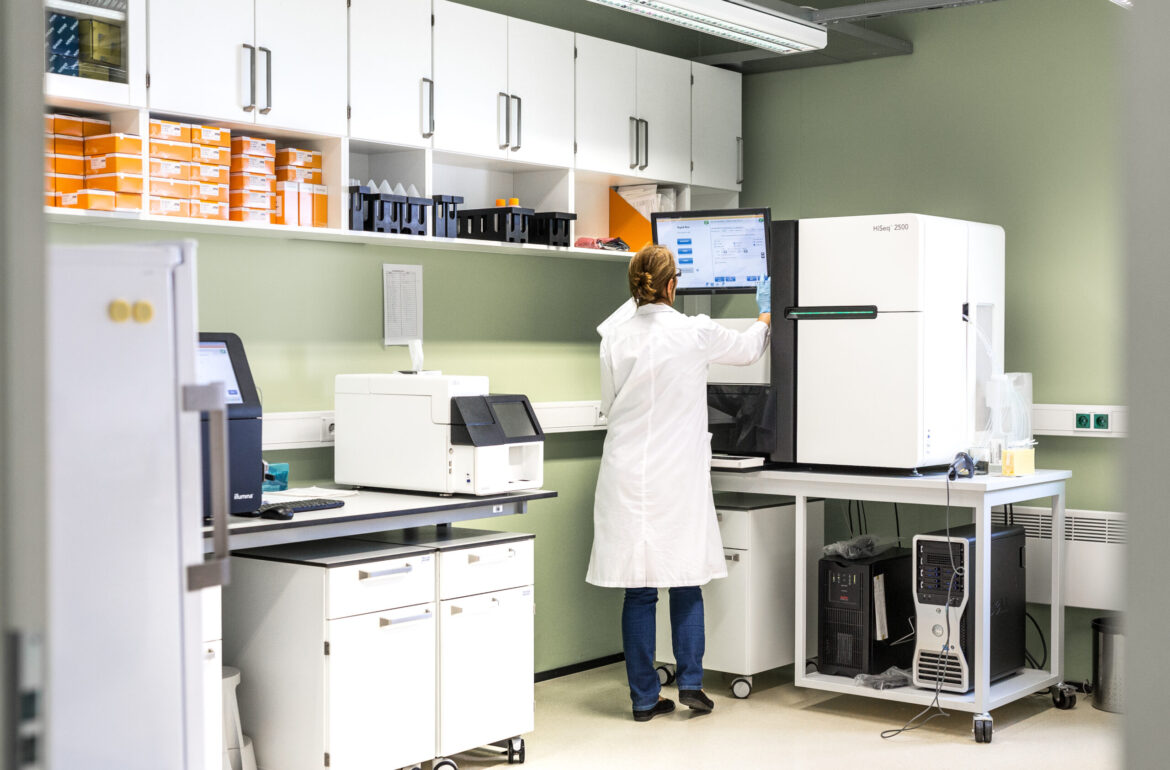
University of Tartu researchers analysed the genomic sequences of coronavirus (SARS-CoV-2) taken from six Estonians. Compared to the initial virus strain that spread in China, researchers have identified eight mutations in Estonia that have been also found in other parts of the world and two mutations which so far seem to be unique to Estonia.
“All the analysed virus strains belong to the A2a clade of SARS-CoV-2, that most probably started to spread in northern Italy and currently causes infection mainly in Europe,” explained Aare Abroi, Specialist in Bioinformatics of the University of Tartu. In addition to mutations that emerged in Europe or elsewhere, two mutations have occurred in the genome of the virus in Estonia.
Further analysis is needed to establish in how many cases the virus has entered Estonia and the extent to which it has spread locally. However, the pattern of the mutations indicates that the Estonian strains can be divided into two groups. “Whether these groups coincide with the outbreaks in Saaremaa and Võru also needs further analysis including epidemiological and clinical data,” said Radko Avi, Senior Research Fellow in Medical Virology of the University of Tartu.
In the first phase of the study, different genomic regions of SARS-CoV-2 were analysed in six virus strains found in samples taken in mid-March. The next goal is to sequence the complete genome of viruses among a larger sample of Estonian coronavirus patients. This would make it possible to identify the paths of the virus and to shed light on cases of infection which have originated from yet unidentified sources.
The long-term goal of the study is to develop a methodology for the analysis of new and yet unknown viral infections, and to generate home-grown knowledge in the field. This would give Estonia the capability to study and diagnose such viral diseases without foreign assistance.
The genomic sequences of the coronavirus strains were determined and analysed by Radko Avi (Senior Research Fellow in Medical Virology of the Institute of Biomedicine and Translational Medicine), Taavi Päll (Research Fellow of Medical Virology from the working group led by Professor of Medical Microbiology Irja Lutsar) and Aare Abroi (Specialist in Bioinformatics of the Institute of Technology). In sequencing, they were supported by Tuuli Reisberg, Manager of Sequencing Core Facility of the Institute of Genomics, Mait Metspalu, Research Professor of Evolutionary Genomics, and Ulvi Gerst Talas, Scientific Programmer of the Institute of Computer Science. Anonymous samples were shared with researchers by the medical laboratory SYNLAB and the study was funded by the University of Tartu.
Further information:
Aare Abroi, Specialist in Bioinformatics, University of Tartu, +372 509 0248, aare.abroi@ut.ee
Radko Avi, Senior Research Fellow in Medical Virology, University of Tartu, +372 5343 3338, radko.avi@ut.ee
The article was originally published on the website of the University of Tartu.
 Back
Back